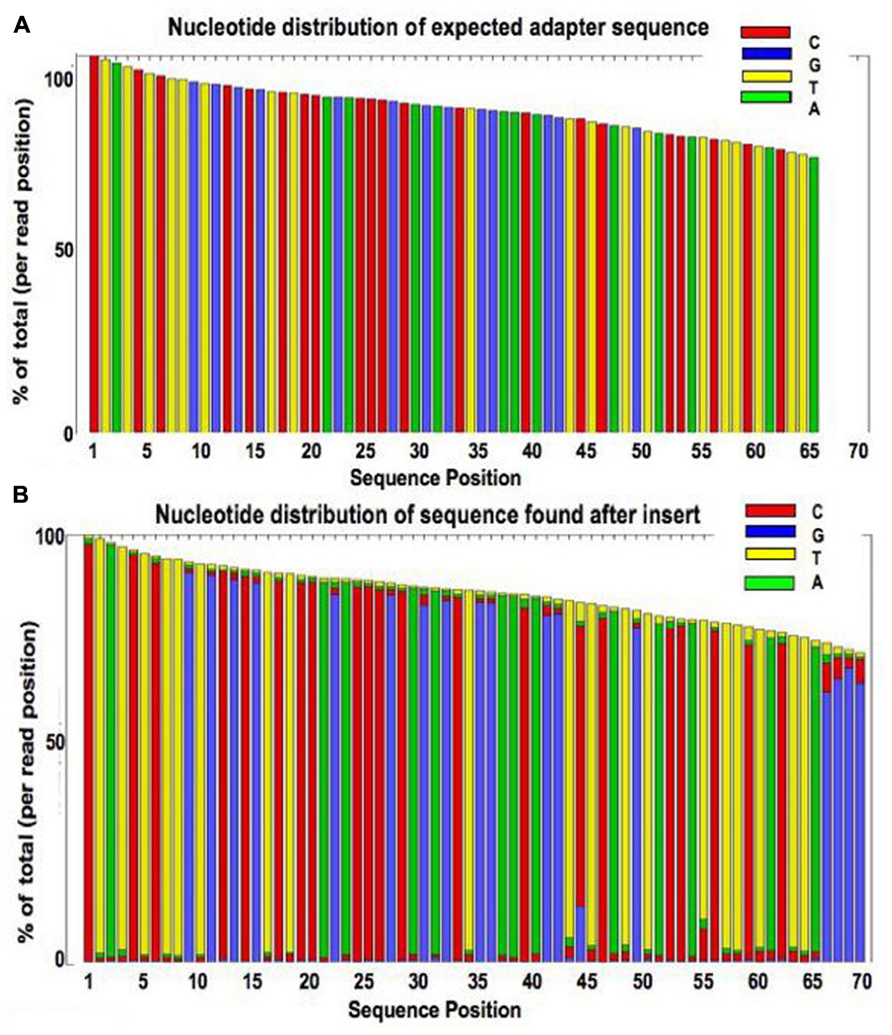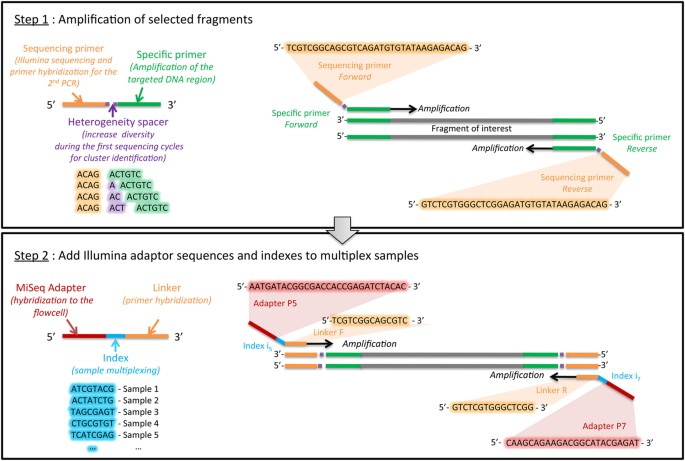
High-throughput sequencing of multiple amplicons for barcoding and integrative taxonomy | Scientific Reports
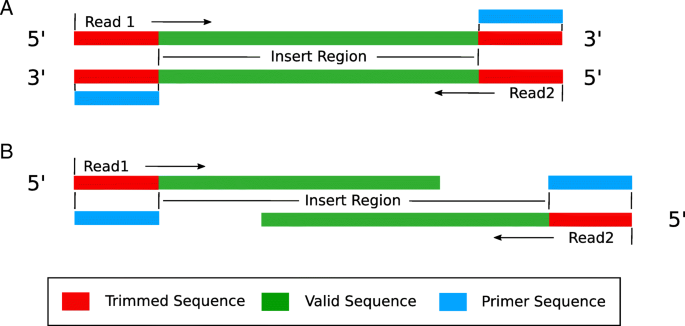
pTrimmer: An efficient tool to trim primers of multiplex deep sequencing data | BMC Bioinformatics | Full Text

AlienTrimmer: a tool to quickly and accurately trim off multiple short contaminant sequences from high-throughput sequencing reads. | Semantic Scholar
Cutadapt remoYes adapter sequences from high-throughput sequencing reads Introduction Implementation Features
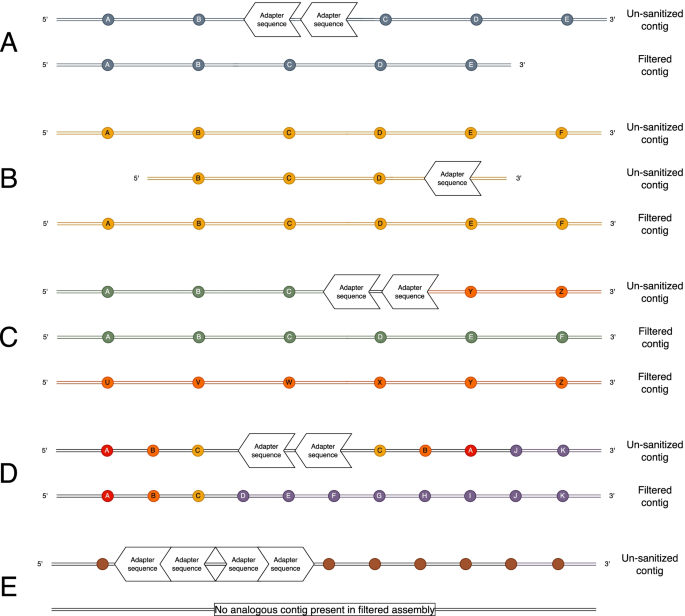
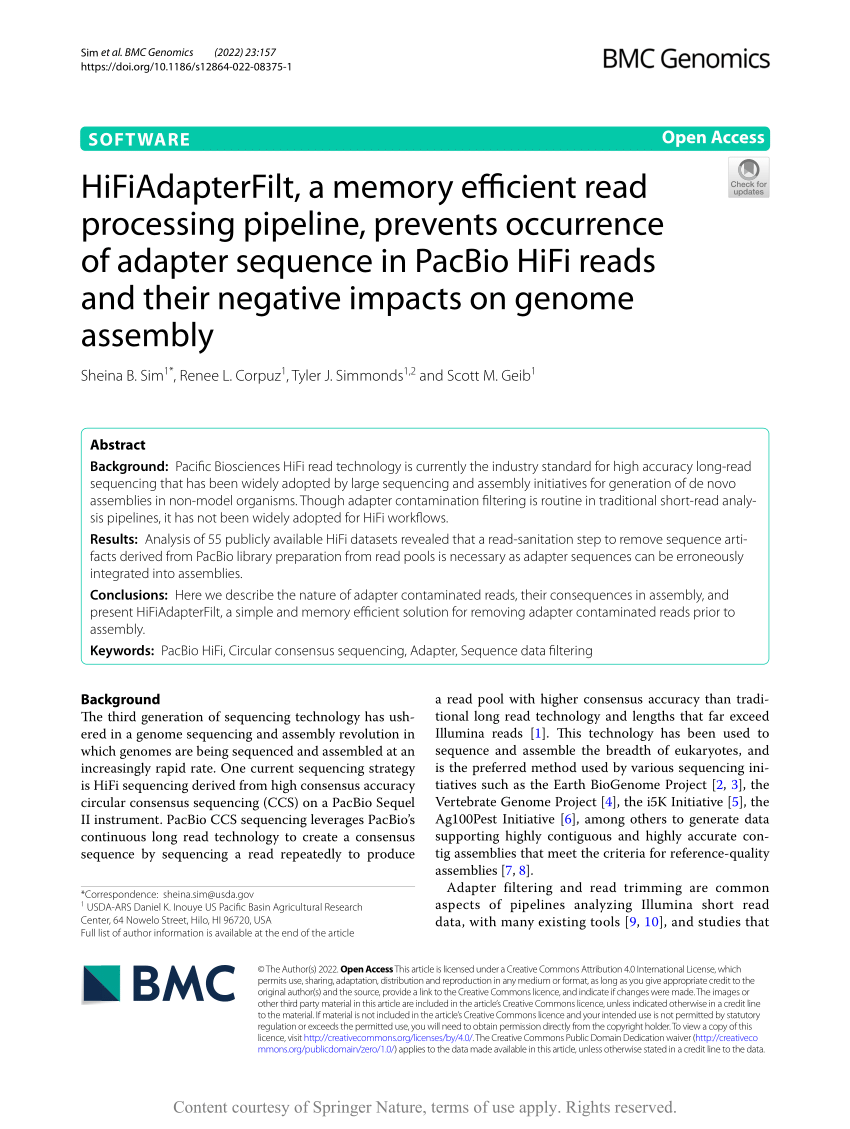
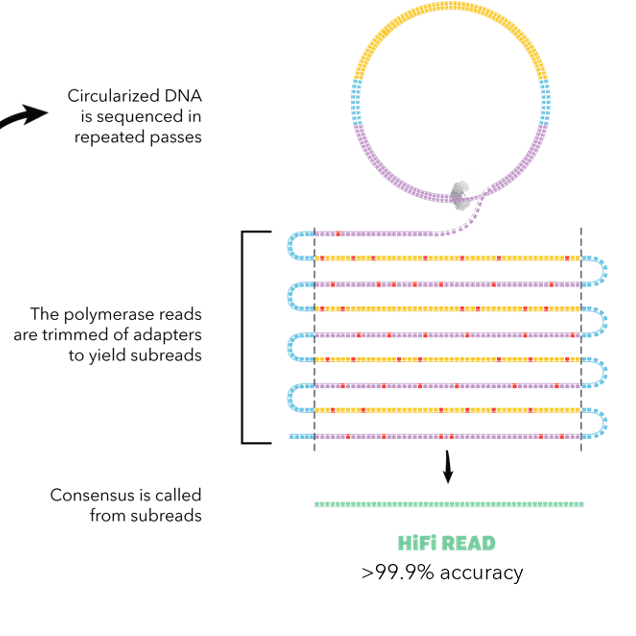
HiFiAdapterFilt, a memory efficient read processing pipeline, prevents occurrence of adapter sequence in PacBio HiFi reads and their negative impacts on genome assembly | BMC Genomics | Full Text

FASTAptamer: A Bioinformatic Toolkit for High-throughput Sequence Analysis of Combinatorial Selections: Molecular Therapy - Nucleic Acids
High-throughput, single-copy sequencing reveals SARS-CoV-2 spike variants coincident with mounting humoral immunity during acute COVID-19 | PLOS Pathogens

cutPrimers: A New Tool for Accurate Cutting of Primers from Reads of Targeted Next Generation Sequencing | Journal of Computational Biology
PathoQC: Computationally Efficient Read Preprocessing and Quality Control for High-Throughput Sequencing Data Sets

An open-sourced bioinformatic pipeline for the processing of Next-Generation Sequencing derived nucleotide reads: Identification and authentication of ancient metagenomic DNA | bioRxiv
From cutadapt to sequencetools (sqt): a versatile toolset for sequencing projects References Relevant Web sites
Comparison of tools cutting primer sequences from reads with tolerance... | Download Scientific Diagram